-Search query
-Search result
Showing 1 - 50 of 2,762 items for (author: cheng & w)

EMDB-37240: 
SARS-CoV-2 Omicron spike in complex with 5817 Fab
Method: single particle / : Cao L, Wang X

EMDB-37241: 
The interface structure of Omicron RBD binding to 5817 Fab
Method: single particle / : Cao L, Wang X

PDB-8khc: 
SARS-CoV-2 Omicron spike in complex with 5817 Fab
Method: single particle / : Cao L, Wang X

PDB-8khd: 
The interface structure of Omicron RBD binding to 5817 Fab
Method: single particle / : Cao L, Wang X

EMDB-37104: 
96-nm axonemal repeat with RS1/2/3
Method: subtomogram averaging / : Cong X, Yao C

EMDB-37111: 
48-nm repeat DMT
Method: subtomogram averaging / : Cong X, Yao C

EMDB-37114: 
Radial Spoke 1 (RS1)
Method: subtomogram averaging / : Cong X, Yao C

EMDB-37116: 
RS1 refined with head mask
Method: subtomogram averaging / : Cong X, Yao C

EMDB-37117: 
Radial Spoke 2 (RS2)
Method: subtomogram averaging / : Cong X, Yao C

EMDB-37118: 
Radial Spoke 2 (RS2) head
Method: subtomogram averaging / : Cong X, Yao C

EMDB-37119: 
Radial Spoke 3
Method: subtomogram averaging / : Cong X, Yao C

EMDB-37120: 
Radial Spoke 3 head
Method: subtomogram averaging / : Cong X, Yao C
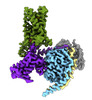
EMDB-43647: 
CryoEM structure of Gi-coupled TAS2R14 with cholesterol and an intracellular tastant
Method: single particle / : Kim Y, Gumpper RH, Roth BL
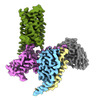
EMDB-43650: 
CryoEM structure of Ggust-coupled TAS2R14 with cholesterol and an intracellular tastant
Method: single particle / : Kim Y, Gumpper RH, Roth BL

EMDB-43656: 
CryoEM structure of Gi-coupled TAS2R14 with cholesterol and an intracellular tastant (Locally refined map)
Method: single particle / : Kim Y, Gumpper RH, Roth BL

EMDB-43657: 
CryoEM structure of Ggust-coupled TAS2R14 with cholesterol and an intracellular tastant (Locally refined map)
Method: single particle / : Kim Y, Gumpper RH, Roth BL

PDB-8vy7: 
CryoEM structure of Gi-coupled TAS2R14 with cholesterol and an intracellular tastant
Method: single particle / : Kim Y, Gumpper RH, Roth BL

PDB-8vy9: 
CryoEM structure of Ggust-coupled TAS2R14 with cholesterol and an intracellular tastant
Method: single particle / : Kim Y, Gumpper RH, Roth BL

EMDB-36005: 
Cryo-EM structure of thehydroxycarboxylic acid receptor 2-Gi protein complex bound MK-6892
Method: single particle / : Yuan Q, Zhu S, Duan J, Xu HE, Duan X

EMDB-36007: 
Cryo-EM structure of thehydroxycarboxylic acid receptor 2-Gi protein complex bound niacin
Method: single particle / : Yuan Q, Zhu S, Duan J, Xu HE, Duan X

PDB-8j6i: 
Cryo-EM structure of thehydroxycarboxylic acid receptor 2-Gi protein complex bound MK-6892
Method: single particle / : Yuan Q, Zhu S, Duan J, Xu HE, Duan X

PDB-8j6l: 
Cryo-EM structure of thehydroxycarboxylic acid receptor 2-Gi protein complex bound niacin
Method: single particle / : Yuan Q, Zhu S, Duan J, Xu HE, Duan X

EMDB-41672: 
ELIC5 with Propylamine in spNW15 nanodiscs with 2:1:1 POPC:POPE:POPG
Method: single particle / : Dalal V, Arcario MJ, Petroff II JT, Deitzen NM, Tan BK, Brannigan G, Cheng WWL

EMDB-41673: 
ELIC with Propylamine in spNW15 nanodiscs with 2:1:1 POPC:POPE:POPG
Method: single particle / : Dalal V, Arcario MJ, Petroff II JT, Deitzen NM, Tan BK, Brannigan G, Cheng WWL

PDB-8twv: 
ELIC5 with Propylamine in spNW15 nanodiscs with 2:1:1 POPC:POPE:POPG
Method: single particle / : Dalal V, Arcario MJ, Petroff II JT, Deitzen NM, Tan BK, Brannigan G, Cheng WWL

PDB-8twz: 
ELIC with Propylamine in spNW15 nanodiscs with 2:1:1 POPC:POPE:POPG
Method: single particle / : Dalal V, Arcario MJ, Petroff II JT, Deitzen NM, Tan BK, Brannigan G, Cheng WWL

EMDB-42977: 
Cryo-EM structure of singly-bound SNF2h-nucleosome complex with SNF2h at inactive SHL2 (conformation 1)
Method: single particle / : Chio US, Palovcak E, Armache JP, Narlikar GJ, Cheng Y

EMDB-43000: 
Cryo-EM structure of SNF2h-nucleosome complex (consensus structure)
Method: single particle / : Chio US, Palovcak E, Armache JP, Narlikar GJ, Cheng Y

EMDB-43001: 
Cryo-EM structure of SNF2h-nucleosome complex (single-bound structure)
Method: single particle / : Chio US, Palovcak E, Armache JP, Narlikar GJ, Cheng Y

EMDB-43002: 
Cryo-EM structure of doubly-bound SNF2h-nucleosome complex
Method: single particle / : Chio US, Palovcak E, Armache JP, Narlikar GJ, Cheng Y

EMDB-43003: 
Cryo-EM structure of singly-bound SNF2h-nucleosome complex with SNF2h at inactive SHL2 (conformation 2)
Method: single particle / : Chio US, Palovcak E, Armache JP, Narlikar GJ, Cheng Y

EMDB-43004: 
Cryo-EM structure of doubly-bound SNF2h-nucleosome complex (conformation 1)
Method: single particle / : Chio US, Palovcak E, Armache JP, Narlikar GJ, Cheng Y

EMDB-43005: 
Cryo-EM structure of doubly-bound SNF2h-nucleosome complex (conformation 2)
Method: single particle / : Chio US, Palovcak E, Armache JP, Narlikar GJ, Cheng Y

PDB-8v4y: 
Cryo-EM structure of singly-bound SNF2h-nucleosome complex with SNF2h at inactive SHL2 (conformation 1)
Method: single particle / : Chio US, Palovcak E, Armache JP, Narlikar GJ, Cheng Y

PDB-8v6v: 
Cryo-EM structure of doubly-bound SNF2h-nucleosome complex
Method: single particle / : Chio US, Palovcak E, Armache JP, Narlikar GJ, Cheng Y

PDB-8v7l: 
Cryo-EM structure of singly-bound SNF2h-nucleosome complex with SNF2h at inactive SHL2 (conformation 2)
Method: single particle / : Chio US, Palovcak E, Armache JP, Narlikar GJ, Cheng Y

EMDB-36634: 
5-HT1A-Gi in complex with compound (R)-IHCH-7179
Method: single particle / : Chen Z, Xu P, Huang S, Xu HE, Wang S

PDB-8jt6: 
5-HT1A-Gi in complex with compound (R)-IHCH-7179
Method: single particle / : Chen Z, Xu P, Huang S, Xu HE, Wang S

EMDB-36541: 
Novel Anti-phage System
Method: single particle / : Li J, Wang Z, Wang L

EMDB-36563: 
Structure of Gabija GajA-GajB 4:4 Complex
Method: single particle / : Li J, Wang Z, Wang L
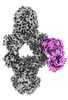
EMDB-36569: 
Novel Anti-phage System
Method: single particle / : Li J, Wang Z, Wang L

EMDB-37915: 
GajA tetramer with ATP
Method: single particle / : Li J, Wang Z, Wang L

EMDB-37916: 
Structure of Gabija GajA in complex with DNA
Method: single particle / : Li J, Wang Z, Wang L

EMDB-38058: 
Cryo-EM structure of Gabija GajA in complex with DNA(focused refinement)
Method: single particle / : Li J, Wang Z, Wang L

EMDB-38070: 
tetramer Gabija with ATP (local refinement)
Method: single particle / : Li J, Wang Z, Wang L

EMDB-38071: 
Tetramer Gabija with ATP
Method: single particle / : Li J, Wang Z, Wang L
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search







 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
